|
|
|
|
|
BlastR
A tool for searching and clustering RNA |
|
|
|
|
|
What is BlastR?
|
BlastR is a new method for searching Non-Coding RNAs in databases. The strategy we adopted relies on the use of the mutual information embedded in di-nucleotides.
We have shown that this approach has better sensitivity and specifity than other softwares with comparable computational cost.
BlastR package is a perl wrapper for BlastP and it is part of the T-Coffee distribution. The BlastR paper is:
BlastR--fast and accurate database searches for non-coding RNAs.
Bussotti G, Raineri E, Erb I, Zytnicki M, Wilm A, Beaudoing E, Bucher P, Notredame C.
Nucleic Acids Res. 2011 May 30 [medline][pdf]
|
|
|
- Download the package here
|
|
|
|
|
|
|
|
|
In order to use blastallR.pl, formatdbR.pl and blastclustR.pl the former installation of the NCBI-Blast package is required.
In order to use blastR.pl and xdformatR.pl the former installation of the AB-Blast package is required.
In order to use wublastR.pl the former intallation of the WU-Blast package is required. WU-Blast is not supported anymore but its upgrade, AB-Blast, is currently distributed.
BlastR package don't require any installation process. Just run the Perl scripts which in turn will call either NCBI-Blast, AB-Blast or WU-Blast on your computer.
|
|
|
|
|
|
|
|
THE PIPELINE
|
Starting from ncRNA alignments we designed a strategy to estimate a Blosum-like scoring matrix that takes into account the cost for aligning two nucleotides while taking into account the nature of their immediate nucleotide neighbors. The strategy consists in converting the RNA sequences (both queries and databeses) into a recoded alphabet (reRNA). This translation consists in attributing to each possible dinucleotide pair an aminoacid-like symbol.This framework make it possible to use BlastP in RNA database search.
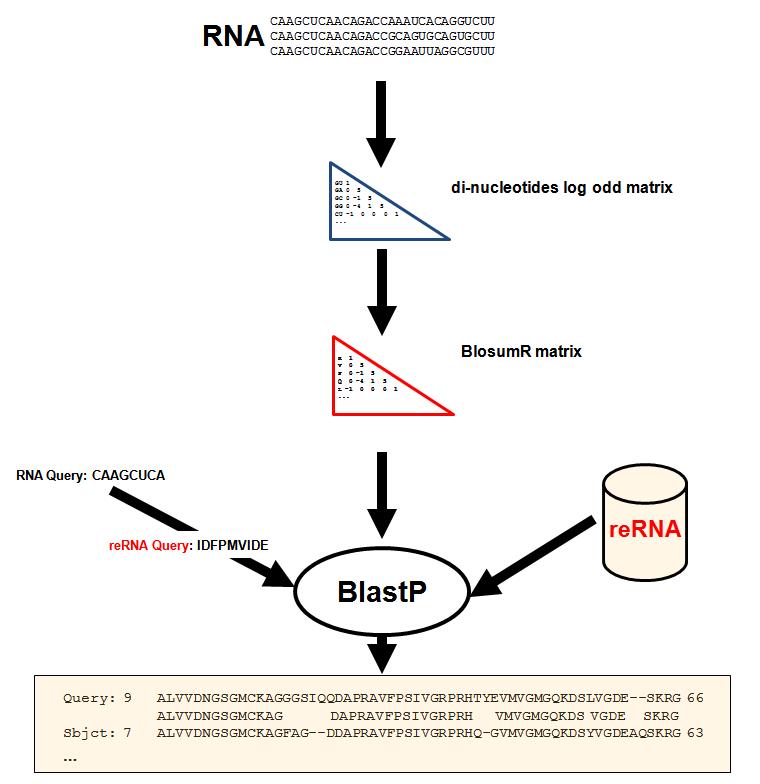
On the left and on the right parts of the figure we show how both query and database (RNA) are converted according to reRNA. On the center of the figure is shown
the BlosumR matrix computation.
|
|
|
|
The scripts provided in blastR package aim to be similar to NCBI-Blast and AB-Blast in terms of usage.
This means that you can pass to blastR scripts the options that normally you would use for NCBI-Blast, AB-Blast or WU-Blast.
The full documentation is available in the download package. But the following shortcuts may be useful.
|
| To format your input multifasta file type |
formatdbR.pl -i <inputMultifasta> -n <databaseOutputName>
|
|
| To run blastallR type |
blastallR.pl -p blastr -i <query> -d <databaseOutputName>
|
|
| To cluster your input multifasta file type |
blastclustR.pl -i <inputMultifasta>
|
|
|
|
|
|
|
|
|
Our projects rely on your feeback. Please send me an
E-mail if you wish to make a request, a comment, or report a bug!
*******************************************
Dr. Cedric Notredame, PhD.
Group Leader
Comparative Bioinformatics Group
Bioinformatics and Genomics Programme
Center for Genomic Regulation (CRG)
Dr Aiguader, 88
08003 Barcelona
Spain
Email: cedric.notredame@gmail.com
HOME : http://www.tcoffee.org/homepage.html
GROUP: CRG
Phone: +34 933 160 271
*******************************************
|
|
|
|
|