|
|
|
|
|
Robusta
A Meta-Multiple Genome Alignment Tool |
|
|
|
|
|
What is Robusta?
|
Robusta is a meta multiple genome alignment package. It does not produce any alignment by itself but can combine alignments of different genome alignment programs into a single final alignment. It is able to call serveral different pairwise and multiple genome alignment programs. The results of these programs are parsed and turned into a T-Coffee library. Using T-Coffee the alignments are then turned into the final alignment.
Currently two pairwise genome aligner and three multiple genome alignment programs are included into this framework.
A typical usage would be to cut the genomes to align into a set of collinear homologous blocks using Mercator. These blocks are than aligned and turned into libraries using Robusta. As a final step T-Coffee is called to produce an alignment for each block.
|
|
|
|
|
|
- Robusta is a seperate program for the Linux operating system and needs several programs to be installed: T-Coffee to combine the libraries and one or serveral genome alignment programs.
- Click here to download Robusta binary
|
|
|
|
|
1- Download the executable or use Git to get the source code of the current version: "git clone https://github.com/cbcrg/robusta.git"
|
|
2- If you downloaded the source code change into the g_coffee directory and type "make mode=release".
|
|
|
|
|
|
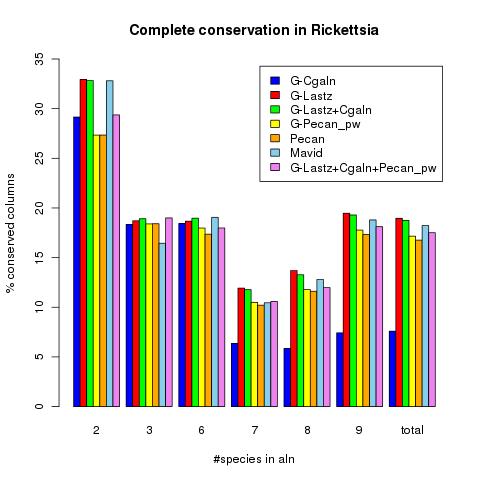
Given a sequence file foo.pep Robusta produces a library which can be turned into an alignment using T-Coffee. See below for a small benchmark using different combinations of genome alignment programs.
9 Rickettsia genomes were first seperated into blocks using Mectator and than aligned using Robusta with different alignments. All Robusta Variants are starting with G.
|
|
|
|
The full documentation is on the T-Coffee Homepage. But the following shortcuts may be useful.
|
| To run Robusta using lastz and cgaln, type |
g_coffee -i foo.seq -p lastz,cgaln -o foo.lib
And then use T-Coffee to produce the final alignment:
t_coffee -lib foo.lib
|
|
| For an overview of all options type: |
g_coffe -h
|
|
|
|
|
|
|
MSA Mehods Available for combination within M-Coffee
|
|
|
|
Our projects relie on your feeback. Please send me an
E-mail if you wish to make a request, a comment, or report a bug!
*******************************************
Dr. Cedric Notredame, PhD.
Group Leader
Comparative Bioinformatics Group
Bioinformatics and Genomics Programme
Center for Genomic Regulation (CRG)
Dr Aiguader, 88
08003 Barcelona
Spain
Email: cedric.notredame@gmail.com
HOME : http://www.tcoffee.org/homepage.html
GROUP: CRG
Phone: +34 933 160 271
*******************************************
|
|
|
|